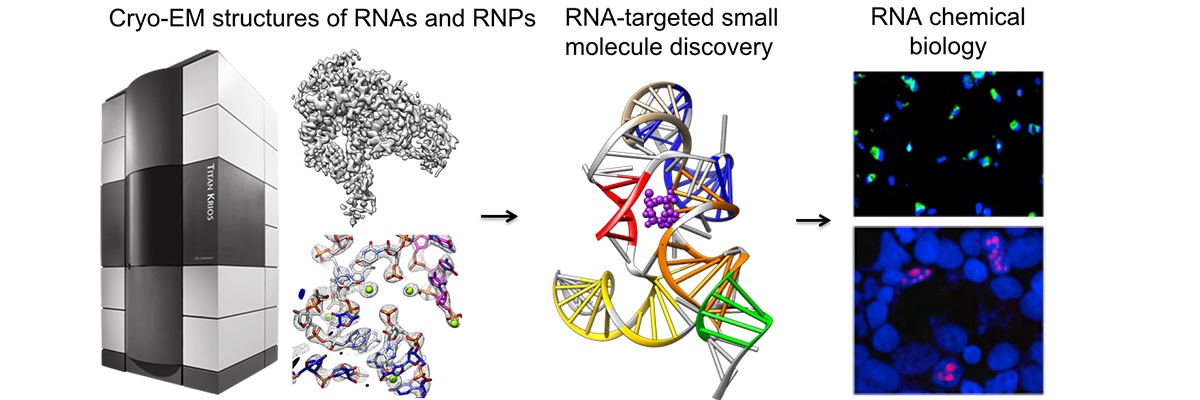
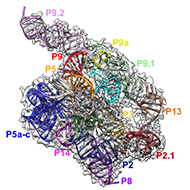
Our lab thrives to use cryo-EM single particle analysis (SPA) to study high resolution structures and functions of RNAs and RNPs when their bound protein partners are involved. We are particular interested in pathogenic non-coding RNAs in bacteria, virus and mammalian systems.
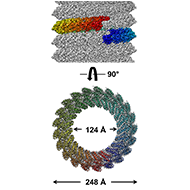
Our lab uses cryo-EM helical reconstruction to understand RNA-binding protein assembly mechanisms of single-stranded RNA virus, which will facilitate development of novel antiviral therapy.
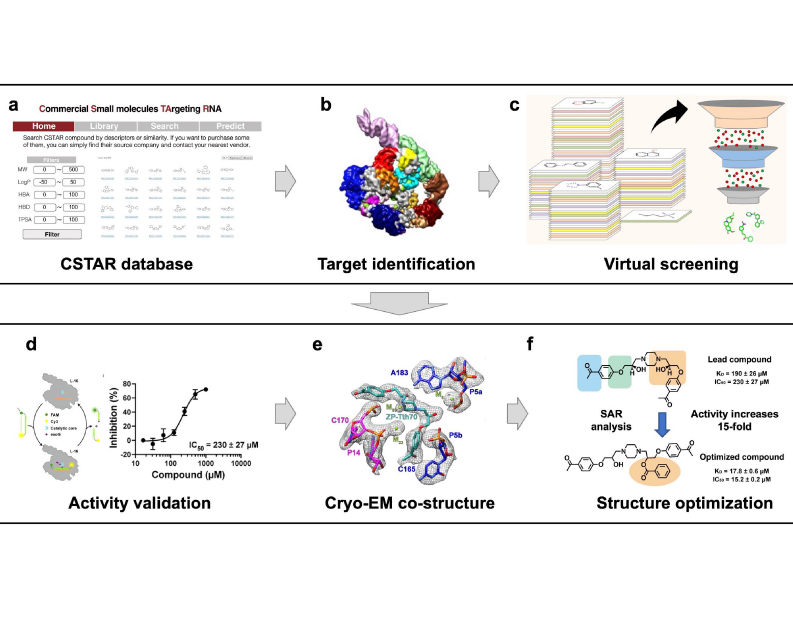
Our lab focuses on understanding RNA structures and developing small-molecule therapeutics targeting RNA. We combine advanced molecular representation learning, virtual screening, and cryo-EM structural analysis to identify and characterize RNA-binding small molecules. Using strategies like HitSTARS, we have built a curated RNA-targeted chemical library (CSTAR) and discovered novel compounds that allosterically regulate RNA function.